Developmental genomics and evolution - A. KHILA
Developmental genomics and evolution

Contact Abderrahman Khila HERE
|
|
Video: presentation of research topics developed by the team of Abderrahman Khila (in french) |
|---|
|
Overview : We are interested in how phenotypes originate and evolve. To do this, we investigate the role of molecular and developmental genetic processes and their interplay with selection in driving phenotypic evolution. We integrate approaches from various fields including EvoDevo, ecology, evolution, genetics, epigenetics, comparative genomics and transcriptomics. As a study model, we ask these questions in water striders – a group of predatory water surface-dwelling insects. Various tools and resources are available to us to study gene expression, gene function, classical genetics and mapping, as well as genomics. |
|
Current projects: There are three primary and interconnected projects ongoing in our lab: |
|
Genetics, epigenetics, and phenotypic plasticity The first deals with the mechanisms underlying the emergence and maintenance of phenotypic variation in a male-specific exaggerated, sexually selected, trait. This project will study the contributions of genetics, epigenetics and nutrition in generating high variation in leg growth in the water strider Microvelia longipes (M. longipes). Males of M. longipes exhibit a spectacular variation in the length of their third legs, whereas females don’t. Behavioural observations, both in the lab and in the wild, showed that males in this species are territorial and use their long legs as weapons during contests to dominate egg-laying sites (Toubiana and Khila, 2019). This extreme variation in males’ leg length is under the influence of both genetic and environmental factors, particularly nutrition (Toubiana and Khila, 2019; Toubiana et al. 2021; Toubiana, Armisen et al. 2021).
|
|
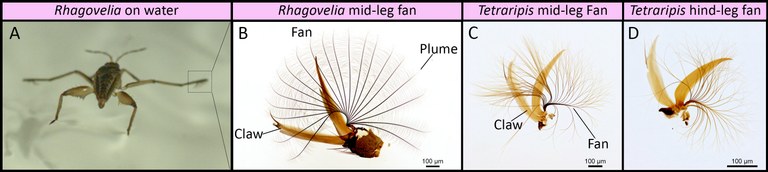
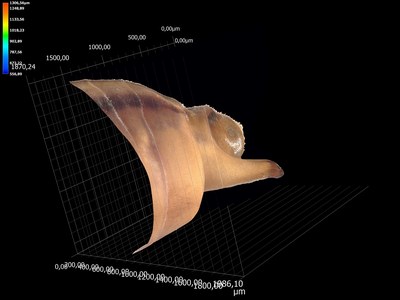
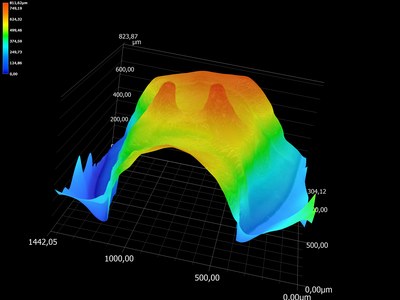
Origin of evolutionary innovations The second investigates the cellular and developmental genetic origin of evolutionary innovations (See Santos et al. 2017). The propelling fan is an evolutionary innovation found in a genus of water striders called Rhagovelia (Andersen, 1982). We found that this evolutionary novelty allowed Rhagovelia to conquer a distinct niche of running waters that is not accessible to closely related species (Santos et al., 2017). We also found that the fan develops during embryogenesis under the control of two taxon-restricted genes that we named geisha (gsha) and mother-of-geisha (mgsha) (Santos et al., 2017). The evolutionary history and the composition of the gene network that led to such an elaborate innovation remain unknown. This project will dissect in more detail the evolutionary steps and the gene network composition underlying the development and evolution of the fan.
|
|
Sexual conflict and co-evolution of the sexes at the genomic level The third project asks how antagonistic co-evolution of the sexes at the phenotypic level is linked to evolution at the genomic level, in the context of sexual conflict over mating rate (see Khila et al. 2012; Crumiere and Khila 2019). Selection in males and females is often antagonistic in that traits favoured in one sex often impose fitness costs to the other. While we have a good understanding of antagonistic co-evolution of the sexes at the phenotypic level, our understanding of how this co-evolution operates at the genomic level remains fragmented at best. This is primarily due to the dichotomy between models where we understand genetics and those where we understand sexual conflict. This project will take advantage of water striders as powerful models for the study of sexual conflict, combined with the genetics and genomics tools our lab has established recently, and resolve this dichotomy. We focus on two genera of water striders where species-pairs, that can be crossed, exhibit prominent differences in a set of male and female sexually antagonistic traits.
|
|
Related publications: * William Toubiana, David Armisén, Corentin Dechaud, Roberto Arbore, and Abderrahman Khila. Impact of trait exaggeration on sex-biased gene expression and genome architecture in a water strider. 2021 BMC Biology. * Toubiana W, Armisén D, Viala S, Decaras A, Khila A (2021) The growth factor BMP11 is required for the development and evolution of a male exaggerated weapon and its associated fighting behavior in a water strider. PLoS Biol 19(5): e3001157. https://doi.org/10.1371/journal.pbio.3001157 * William Toubiana and Abderrahman Khila. Fluctuating selection strength and intense male competition underlie variation and exaggeration of a water strider's male weapon. Proceedings of the Royal Society B, 2019. * Antonin Jean Johan Crumiere, Abderrahman Khila. Hox genes mediate the escalation of sexually antagonistic traits in water striders. Biology letters 2019. * Santos, M.E., Le Bouquin, A., Crumiere, A.J.J., Khila, A. Taxon-restricted genes at the origin of a novel trait allowing access to a new environment. Science, 2017. * Khila, A., Abouheif, E., Rowe, L. Function, developmental genetics, and fitness consequences of a sexually antagonistic trait. Science, 2012. |
Research Opportunities:
Want to work with us? write an e-mail to Abderrahman Khila
Team members
Past members
| L3 student | ||
| PhD student | ||
| AI, EPHE student | ||
| post-Doc | ||
| IE, CDD CNRS | ||
| IE | ||
| TCH, CNRS |
|
|
| post-doc | ||
| PhD student | ||
| PhD student |
Contact: abderrahman.khila@ens-lyon.fr
Past members
| Full Name | Status | Institution |
|---|---|---|
| ROUX Pascale | TCE | INRAE |
| ARBORE Roberto | Post doc | CNRS |
| ARIAS Leticia | IE CDD | CNRS |
| ARMISEN David | Researcher CDD | ENS de Lyon |
| BERNARD Marie | TECH | CNRS |
| BONNETON François | MCF HC | UCBL |
| BOS Celeste | Stagiaire | ENS de Lyon |
| CRUMIERE Antonin | PhD student | CNRS |
| DECARAS Amélie | AI, EPHE student | CNRS |
| DUPUIS Benjamin | Licence Student | ENS |
| FINET Cédric | post-Doc | CNRS |
| LE BOUQUIN Augustin | PhD Student | - |
| MIGNEREY Ilona | Stagiaire M2 | ENS de Lyon |
| MONTBEL Vincent | M2 student | INSA |
| ROUSSEL Alice | M2 Student | UCBL |
| RUTKOWSKA Maria | Stagiaire L | ENS de Lyon |
| SAKER Kahina | DUT Student | CNRS |
| SANTOS Emilia | post-Doc- | - |
| SZENTKIRALYI Fruzsina | Stagiaire M2 | ENS de Lyon |
| TOBIAS SANTOS Vitoria | PhD student | CNRS |
| TOUBIANA William | PhD student | ENS de Lyon |
| VARGAS-LOWMAN Aïdamalia | PhD student | - |